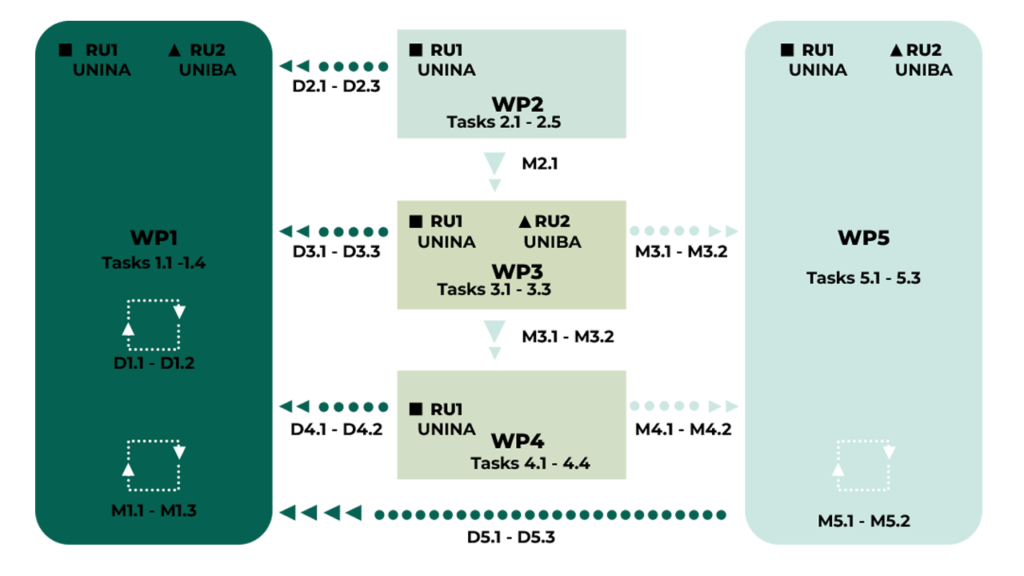
The project activities are structured into 5 interconnected Work Packages (WPs) carried out over two years.
WP1: Scientific Coordination and Management (RU1, RU2): Management of the project, internal progress reports, coordination of activities, and organization of meetings.
WP2: Production of Fungal SMs (RU1): Selection of fungal strains (Talaromyces pinophilum and Diplodia corticola) and optimization of growth conditions for the production and chemical identification of secondary metabolites
WP3: Evaluation of Antiviral Activity (RU1, RU2): Determination of the maximum non-cytotoxic dose (IC50) and in vitro evaluation of antiviral activity of SMs against CCoV and FCoV.
WP4: Mechanism of Action (RU1): In vitro and in silico investigations into the mechanism of action, focusing on the role of AhR and FPRs in virus-host interaction, and chelation studies with metal ions (iron).
WP5: Dissemination and Exploitation (RU1, RU2): Raising awareness, publishing results in Open Access journals, participating in conferences, and promoting project outcomes
Detailed Description of the Project
Methodologies, Objectives, and Expected Results
The project involves two Research Units (RUs), coordinated by RU1, and articulated over two years into 5 interconnected Work Packages (WPs). Each WP includes specific tasks (Ts), deliverables (Ds), and milestones (Ms), ensuring coherence with strategic themes and innovative objectives.
The project highlights sector relevance, positioning within the state of the art, and details the contribution of each research unit.
WP1: Scientific Coordination and Project Management (RU1, RU2)
Objectives
The WP1 includes the organization and overall management of the research proposal organizing all the activities which will allow for the project advancement. To share information and solve problems and conflicts, a constant communication flow within the RUs will be guaranteed.
PI, certifying the project activities, evaluating results, deliverables, and milestones, WP progress, will performed bimonthly reports on progress in achieving of both intermediate and final milestones and targets. Unit responsible, bimonthly, will report on sustained expenses incurred and will send to MUR.
Methodology
- T1.1. – Organization of the kick-off meeting.
- T1.2. – Coordination of the scheduled activities of the project. This task coordinates the management of the research proposal and the evaluation of results, through internal reports and subsequent feedback actions. To develop the common objectives as well as to address potential emerging critical issues, the scientific competence and the availability of the RUs components will be basic.
- T1.3. – Organization of meetings. To have discussions among the RUs about works progression, three or four web-meetings/year will be scheduled. Moreover, at least one physical meeting/year will be also scheduled.
- T1.4. – Prepare financial reports. Administrative offices will provide the reporting of each RU and will prepare the financial report (intermediate and final).
Deliverables
- D1.1 Procedures for administrative management
- D1.2 Procedures for delivery of internal and progress reports
Milestones
- M1.1 Kick-off and periodic internal progress meeting;
- M1.2 Bimonthly scientific and financial reports;
- M1.3 Final conference.
WP2: Production of Fungal SMs (RU1)
Objectives
Selection of strains of Talaromyces pinophilum and Diplodia corticola with purity and quantity suitable for project activities.
Methodology
- T2.1. Selection of strains of Talaromyces pinophilum. The isolates, already available in the collections present in the proposers’ laboratories, will be selected based on their ability to produce funicones as OMF and vermistatin. In particular, liquid cultures of isolates were prepared by inoculating mycelial plugs from actively growing cultures of selected strains in 500 mL-Erlenmayer flasks containing 250 mL potato–dextrose broth (PDB) which were kept in darkness on stationary phase at 25 °C. After 21 days, cultures were filtered at 0.45 μm and subjected to LC-MS analysis. Known funicones will be identified by comparison with the data reported in a custom LC-MS library.
- T2.2. Selection of strains of Diplodia corticola. The isolates, already available in the collections present in the proposers’ laboratories, will be selected based on their ability to produce sphaeropsidines (Sphs) as SphA-C. Liquid cultures of the strain were prepared in Czapek-Dox broth amended with 2% corn meal in 500 mL Erlenmeyer flasks containing 250 mL of the substrate and grown on stationary phase in the dark at 25°C. After 30 days, cultures were filtered at 0.45 μm and subjected to LC-MS analysis. Known Sphs will be identified by comparison with the data reported in a custom LC-MS library.
- T2.3. Optimization of growth conditions. For this purpose, the strains identified in the T1.1/T1.2 will be grown in liquid cultures on two different culture media in reduced aliquots at specific times and culture conditions. Fungal culture filtrates (CFs) will be subjected to extraction with suitable organic solvent to obtain crude extracts. These latter will be subjected to qualitative and quantitative analyses by using a GC- or LC MS systems.
- T2.4. Production and extraction of bioactive metabolites. The CFs, produced in large amount according to the growth conditions selected in the Task 1.2, will be subjected to extraction with organic solvents, while the purification of crude extracts will be obtained by suitable chromatographic methods (essentially adsorption chromatography on column and/or TLC of silica gel with eluent mixtures with different polarity) according with protocols previously developed by proponents.
- T2.5. Identification of secondary metabolites. This activity will be carried out through comparison of spectroscopic (NMR) and mass data with those reported in the literature for funicones and sphaeropsidins. The structure elucidation of new metabolites will be performed using spectroscopic methods (IR, UV, mono and bi-dimensional proton, and carbon NMR (nuclear magnetic resonance), EI (electron ionization), and using chemical methods which, through functional group transformation, will allow us to confirm structures.
Deliverables
- D2.1 Report on identification of best cultivation conditions
- D2.2 Report on best producer strains
- D2.3 Report on structure elucidation of known and new SMs
Milestones
- M2.1 Availability of SMs
WP3: Identification of IC50, Dose-Response Curves, and In Vitro Antiviral Testing (RU1, RU2)
Objectives
The aim of WP3 will be the determination of in vitro cytotoxicity on non-infected cells of SMs selected and produced in WP2. The in vitro evaluation of antiviral activity against CCoV and FCoV of the SMs obtained by previous Task will be investigated. Contextually, the action of these compounds at different steps in the viral infection cycle will be described.
Methodology
- T3.1. (RU-1) Developing dose-response curve of SMs. Canine fibrosarcoma (A72) cells and Crandell-Rees Feline Kidney (CRFK) cells will be grown in monolayer culture with Dulbecco Minimal Essential Medium (DMEM), supplemented with 10% fetal bovine serum (FBS), 100 μg/mL penicillin and 100μg/mL streptomycin. Cells will be incubated at 37°C in a 5% CO2 incubator (Cerracchio et al., 2022a; Camero et al., 2022). Adherent subconfluent cell monolayers will be prepared in growth medium in 96 well plates for cytotoxicity tests. Cell viability by Trypan blue exclusion test will be first assessed (Fiorito et al., 2022), cell proliferation by MTT will be then performed (Fiorito et al., 2022). Statistical analysis of results will be presented as mean ± S.D. One-way ANOVA with Tukey’s post-test will be performed by GraphPad InStat Version 3.00 for Windows 95 (GraphPad Software, San Diego, CA, USA). p < 0.05 will be considered statistically significant.
- T3.2. (RU1-RU2) – In vitro evaluation of antiviral activity of SMs on CCoV and FCoV infection. A72 cells and CRFK cells will be grown in monolayer culture with DMEM supplemented with 10% FCS, 100 IU/mL penicillin and 0,1 IU/mL streptomycin. Cells will be incubated at 37°C in a 5% CO2 incubator. Cells will be plated into 24-wells culture plates for antiviral assays. The reference strains CCoV S/378 and FCoV 79/1683 will be employed for all the experiments. Infectivity (viruses titres) will be determined by titration assay (Cerracchio et al., 2022a; Camero et al., 2022) and stock virus will be stored at -80°C. Based on the results of the cytotoxicity test, the antiviral activities against CCoV and FCoV will be evaluated using the SMs at the maximum non-cytotoxic dose and below the cytotoxic threshold. To identifying the viral inhibition phase of the selected compounds against CCoV and FCoV, different protocols (A, B, C and D) will be performed. All experiments will be performed in triplicate. Protocol A: virus infection of cell monolayers before treatment with SMs. Protocol B: virus infection of cell monolayers after treatment with SMs. Protocol C: virus infection and in parallel SMs treatment of cell monolayers. Protocol D: virucidal activity.
- T3.3. (RU1-RU2) – Immunofluorescence staining and virus titration assay. Immunofluorescence staining will be performed in vitro with monoclonal antibodies to recognize CCoV and FCoV, at different steps in the viral infection cycle. Contextually, virus titration assays will be performed to evaluate antiviral activity on CCoV and FCoV before and after SMs treatment.
Deliverables
- D3.1 Report on maximum non-cytotoxic dose
- D3.2 Report on antiviral activity
- D3.3 Report on action of SMs during viral infection cycle
Milestones
- M3.1 Identification of IC50
- M3.2 Identification of SMs with antiviral activity
WP4: Mechanism of Action of Effective SMs – In Vitro and In Silico Studies (RU1)
Objectives
To explore the mechanism of action of effective SMs in virus-host interaction, in vitro investigations studies will be performed. Moreover, the determination of stability constants of the complexes of funicones and sphaeropsins and iron will be carried out. Methodology
Methodology
- T4.1. (RU1) – In vitro mechanism of action on AhR and FPRs. To explore the mechanism of action of effective SMs in the potential antiviral activities against CCoV and FCoV, the role of AhR, a receptor expressed in A72 cells, will be investigated through immunofluorescence assay for AhR (Fiorito et al., 2022; Cerracchio et al., 2022a,b). A72 cells will be either infected or not with CCoV and treated with effective SMs. Similarly, CRFK cells will be either infected or not with FCoV and treated with effective SMs. At different times of infection, immunofluorescence staining will be performed using anti-AhR a specific antibody (Fiorito et al., 2022; Cerracchio et al., 2022a,b). Microscopy and photography were assessed by the ZOE Fluorescent Cell Imager (Bio-Rad Laboratories, Hercules, CA, USA). Quantification of fluorescence signals from microscopy-generated images will be performed using ImageJ (National Institutes of Health, Bethesda, MD, USA) software.
The project also aims to explore the role of FPRs in CCoV and FCoV infection and its potential as an innovative therapeutic target. For this purpose, the presence of FPRs in A-72 and CRFK cells will be investigated by western blot and immunofluorescence analyses, using anti-FPRs specific antibodies. Then, A-72 and CRFK cells will be infected with CCoV and FCoV, respectively, in the presence or absence of SMs treatment. The effects of these metabolites in modulating the FPRs activation will be examined. In detail, we will explore: Reactive Oxygen Species (ROS) generation and NADPH oxidase activation by dihydrorhodamine and NADPH oxidase activity test; intracellular calcium influx by colorimetric assay; inflammatory response (analysis of pro and anti-inflammatory cytokines) by bio-plex assay and RT-PCR study (Cuomo et al., 2021); cell migration by in vitro scratch assay and the FPRs downstream signaling pathway by western blot analysis. - T4.2. (RU1) – In vitro mechanism of action on non-heme iron. The ability of some SMs to chelate metal ions such as iron, during CCoV and FCoV infections, will be also preliminarily examined by the classic colorimetric method Perl’s iron staining. Indeed, non-heme iron, including soluble free iron and iron-protein deposits (ferritin and hemosiderin) can be visualized in cells, where soluble ferrocyanide reacts with the tissue Fe3+ to form crystals making an insoluble Prussian blue dye (Meguro et al., 2007). A72 and CRFK cells, exposed or not to effective SMs, will be infected with CCoV and FCoV, respectively. At different times after infection, cells will be washed with PBS, fixed with 95% ethanol, drained and dried. Afterward, cells will be stained with Prussian blue Perls’ staining as previously reported (Fiorito et al., 2021).
- T4.3. (RU1) Chelation studies. The ability of active funicones and Sphs to chelate the iron will be carried out in solution through the use of different techniques such as: potentiometry, UV-Vis spectroscopy, circular dichroism (CD), 1H-NMR spectroscopy and LC-ESI-HRMS. The processing of the experimental data will be carried out through numerical methods of minimization (Gans at al, 1996). By use of these techniques it will be able to characterize the complexation between Fe(II) and Fe(III) and tested compounds. Due to the low solubility of metabolites in water, the stoichiometry and complex formation constants were evaluated in CH3OH/0.1 M NaClO4 (50/50 w/w) ionic medium.
- T4.4. (RU1) – In silico studies of mechanism of action on AhR and FPRs.
In order to further evaluate the AhR as well as FPRs recognition mechanism by effective SMs we will apply a combined strategy in which Molecular Docking calculations will be integrated with homology modelling methodologies.
First, the three-dimensional structure of the AhR and FPR receptors will be predicted from the amino-acid sequence using AlphaFold approach (Jumper et al., 2021) or Protein Data Bank (RCSB, PDB), while the 3D conformation of the selected metabolites will be obtained from PubChem (ncbi library). Successively, the obtained AhR and FPRs three-dimensional structural model will be used as reference structure for the receptors in the following molecular docking studies. Then, all docking process will be performed using AutoDock 4.0 (Morris et al., 2009). The molecular docking protocol will involve the following step: i) preparing the starting coordinate files for the effective SMs and the receptors in order to include the information required (i.e. spatial charges, polar hydrogen atoms, atom types and torsional degrees of freedom). ii) AutoGrid routine: the software AutoDock requires pre-calculated grid maps that are defined by AutoGrid. iii) Docking calculation by AutoDock routine: the input parameters for docking calculation will be contained in a specific file created using the AutoDockTools. In details, the Lamarckian Generic Algorithm (LGA) will be used as searching methods. Minimized effective SMs will be randomly positioned inside the grid box. After docking calculation, the obtained structures will be clustered and will be analysed using Pymol (DeLano, 2002) and Chimera (Pettersen et al., 2004) software.
Deliverables
- D4.1 Report on AhR and FPRs involvement in viral infection
- D4.2 Report on iron-chelation properties of SMs
Milestones
- M4.1 Identification of novel antiviral mechanisms
- M4.2 Identification of new antiviral targets
WP5: Dissemination, Communication, and Exploitation (RU1, RU2)
Objectives
Raise awareness the academic community, researchers, and pharmaceutical companies in discovery of new antivirals and guarantee the transfer, dissemination and exploitation of significant data for further applications. Publications and dissemination.
The new knowledge will be shared with the academic community through publishing in high impact ISI peer-reviewed journals will be published, guaranteeing open access, thus, results will be freely available to both subscribers and the wider public, in agreement with Horizon 2020/Europe policy.
Attention will be paid to intellectual property in case of patenting potentialities. Moreover, we will disseminate our findings through presentations at conferences and seminars, as well as graduate teaching and general public. The results of this project will be also divulgated in national and international conferences. Effective internal communication between RUs components will be ensured through the usage of specific web platforms (e.g., Microsoft Teams, Zoom, etc).
Methodology
- T5.1. Improvement of project audience through the development of a website for the public exchange of the obtained results, containing an overview of the project and main updates. This website will include a captivating logo, an overview of the project, aims, and main activities; social media pages (e.g., Facebook, Twitter, Instagram) will be activated and promptly updated to ensure high visibility.
- T5.2. Conferences and workshops. All the RUs will provide disclosure of the results and raw data. The investigators will organize/participate to National/International conferences to present results in order to gain visibility from the scientific community. T5.3. Papers and patents. The investigators will publish their results in Open Access. The results that will emerge for scientific and/or technological impact will be considered for patenting.
Deliverables
- D5.1 Project website and public communication tools
- D5.2 Peer-reviewed Open Access scientific publications
- D5.3 Intellectual property agreements
Milestones
- M5.1 Development of potential antivirals against CCoV and FCoV
- M5.2 Advancement in understanding viral host–cell interaction mechanisms
